·Phi Coefficient of Association
·Chi-Square Test of Association
·Fisher Exact Probability Test
For a table of frequency data cross-
- calculate the Phi coefficient of association;T
- perform a chi-square test of association, if the sample size is not too small; andT
- perform the Fisher exact probability test, if the sample size is not too large. [Although the Fisher test is traditionally used with relatively small samples, the programming for this page will handle fairly large samples, up to about
n=1000, depending on how the frequencies are arrayed within the four cells.]T
For intermediate values of n, the chi-square and Fisher tests will both be performed.
Data EntryT
| X |
Expected Cell Frequencies per Null Hypothesis
| 0 |
1 |
Totals |
| Y |
1 |
0 |
| Totals |
| | |||||||||
| Chi-Square |
Chi-square is calculated only if all expected cell frequencies are equal to or greater than 5. The Yates value is corrected for continuity; the Pearson value is not. Both probability estimates are non-directional. Phi |
Yates |
Pearson | | |||
Fisher Exact Probability Test:T
| P | one-tailed | |
| two-tailed |
| Home | Click this link only if you did not arrive here via the VassarStats main page. |
©Richard Lowry 2001-
All rights reserved.
Fisher Exact Probability Test: Logic and Procedure
Consider a 2x2 contingency table of the sort described above, with the cell frequencies represented by a, b, c, d, and the marginal totals represented by a+b, c+d, a+c, b+d, and n.
| 0 | 1 | Totals
| 1
| a
| b
| a+b
| 0
| c
| d
| c+d
| Totals
| a+c
| b+d
| n
| |
If there were no systematic association between the variables A and B within the population from which the cell frequencies are randomly drawn, the probability of any particular possible array of cell frequencies, a, b, c, d, given fixed values for the marginal totals a+b, c+d, etc., would be given by the hypergeometric rule
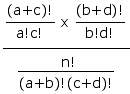
which for computational purposes reduces to
![]()
Also, the degree of disproportion within any array of cell frequencies—
![]()
For any particular observed array of cell frequencies, the programming embedded in this page calculates the probability of that particular array plus the probabilities of all other possible arrays whose degree of disproportion is equal to or greater than that of the observed array. Thus, for the observed array
2
7
9
8
2
10
10
9
19
the one-tailed probability would be the sum of the separate probabilities for the arrays
probability
2
7
8
2
0.01754
1
8
9
1
0.00097
0
9
10
0
0.00001
sum = 0.01852
(one-tailed probability)
And the two-tailed probability would be that sum plus the sum of the separate probabilities for the arrays of equal or greater disproportion at the other extreme:
probability
8
1
2
8
0.00438
9
0
1
9
0.00011
sum = 0.00449
two-tailed probability = 0.01852 + 0.00449 = 0.02301
Return to Top
©Richard Lowry 1998-
All rights reserved.